Abstract
Statistical network models have become popular tools for analyzing multivariate psychological data. In empirical practice, network parameters are often interpreted as reflecting causal relationships – an approach that can be characterized as a form of causal discovery. Recent research has shown that undirected network models are likely to perform poorly as causal discovery tools in the context of discovering acyclic causal structures, a task for which many alternative methods are available. However, acyclic causal models are likely unsuitable for many psychological phenomena, such as psychopathologies, which are often characterized by cycles or feedback loop relationships between symptoms. A number of cyclic causal discovery methods have been developed, largely in the computer science literature, but they are not as well studied or widely applied in empirical practice. In this paper, we provide an accessible introduction to the basics of cyclic causal discovery for empirical researchers. We examine three different cyclic causal discovery methods and investigate their performance in typical psychological research contexts by means of a simulation study. We also demonstrate the practical applicability of these methods using an empirical example and conclude the paper with a discussion of how the insights we gain from cyclic causal discovery relate to statistical network analysis.Key Takeaways
- Psychological research should move beyond assuming simple acyclic cause-and-effect and instead consider cyclic causal models, which account for the feedback loops and reciprocal relationships often present in psychological phenomena.
- This paper reviews and evaluates state-of-the-art methods for causal discovery from purely observational data, a critical task when experimental manipulation is not possible.
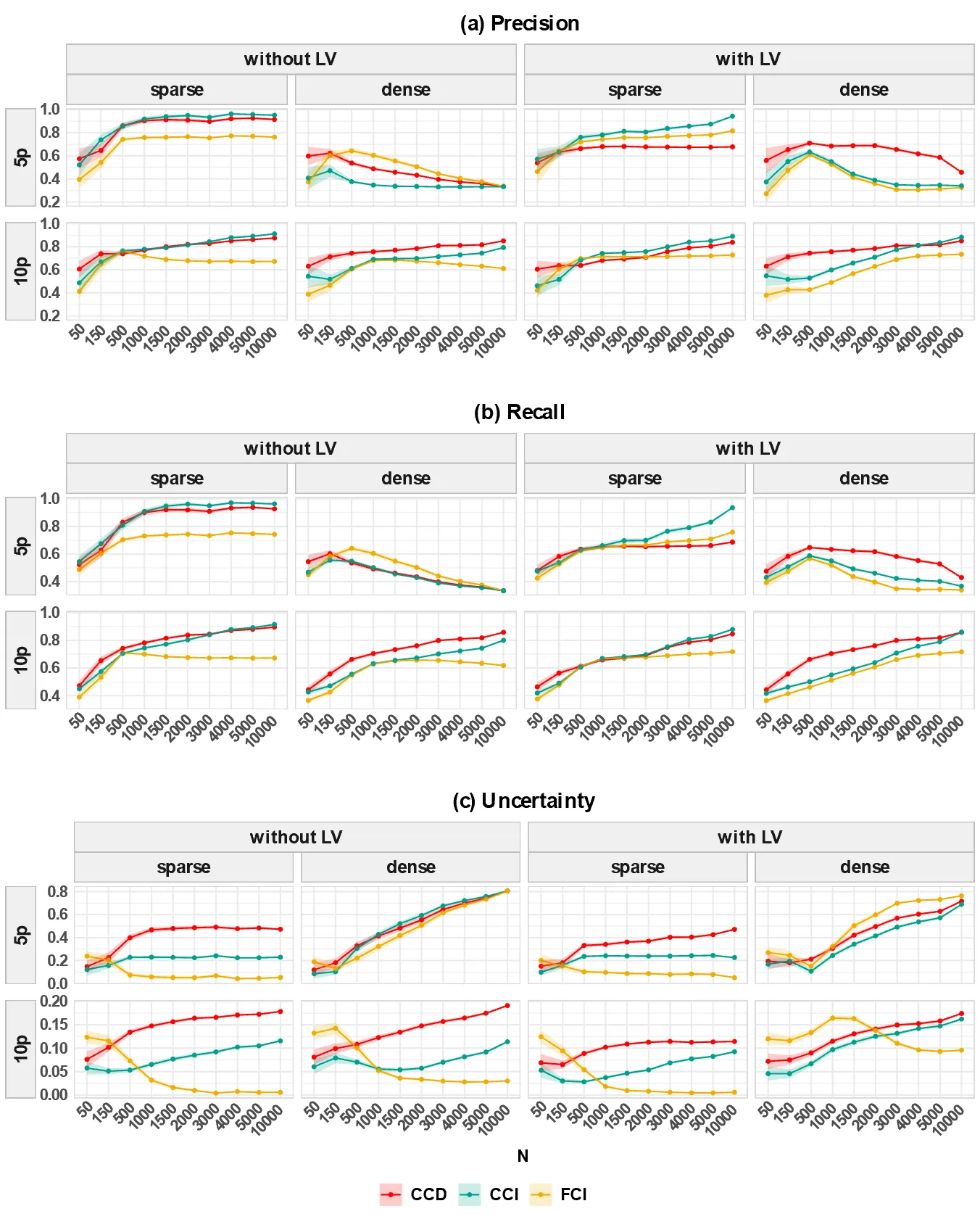
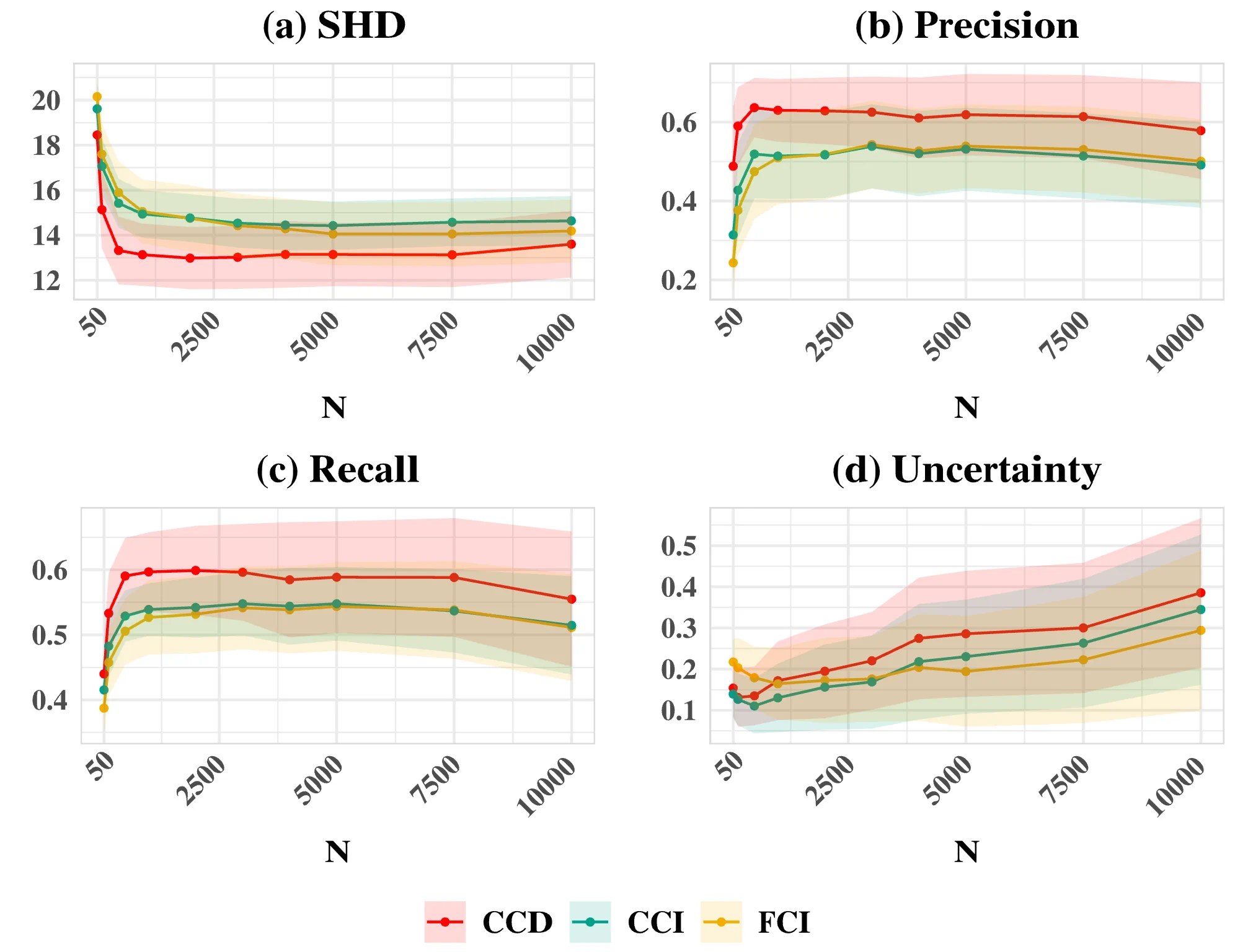
- Simulation studies demonstrate that an autoregressive-based (AR-based) method is the most promising approach, outperforming FCI-variants and LV-based methods in its ability to accurately discover cyclic causal structures from psychologically plausible data.
Author Details
Citation
Park, K., Waldorp, L.J., & Ryan, O. (2024). Discovering cyclic causal models in psychological research. advances.in/psychology, 2, e72425. https://doi.org/10.56296/aip00012
Transparent Peer Review
The current article passed two rounds of double-blind peer review. The anonymous review report can be found here.







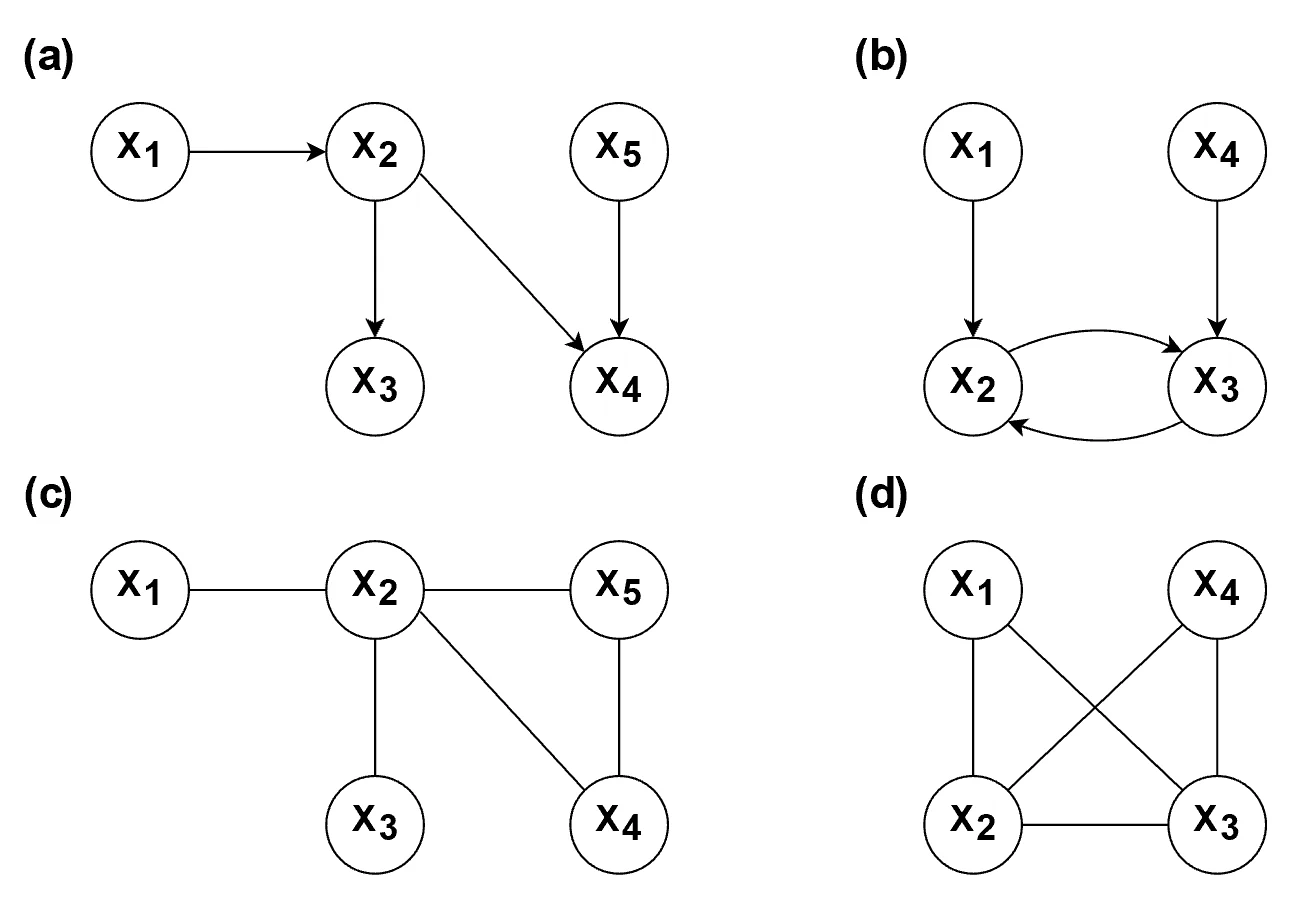
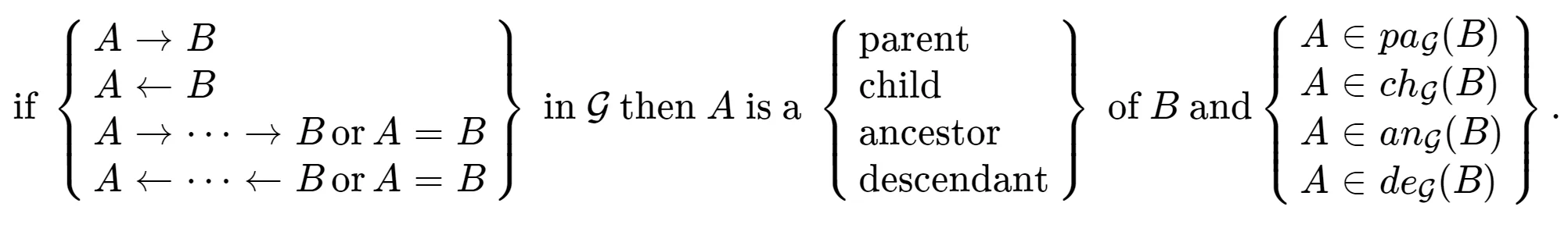
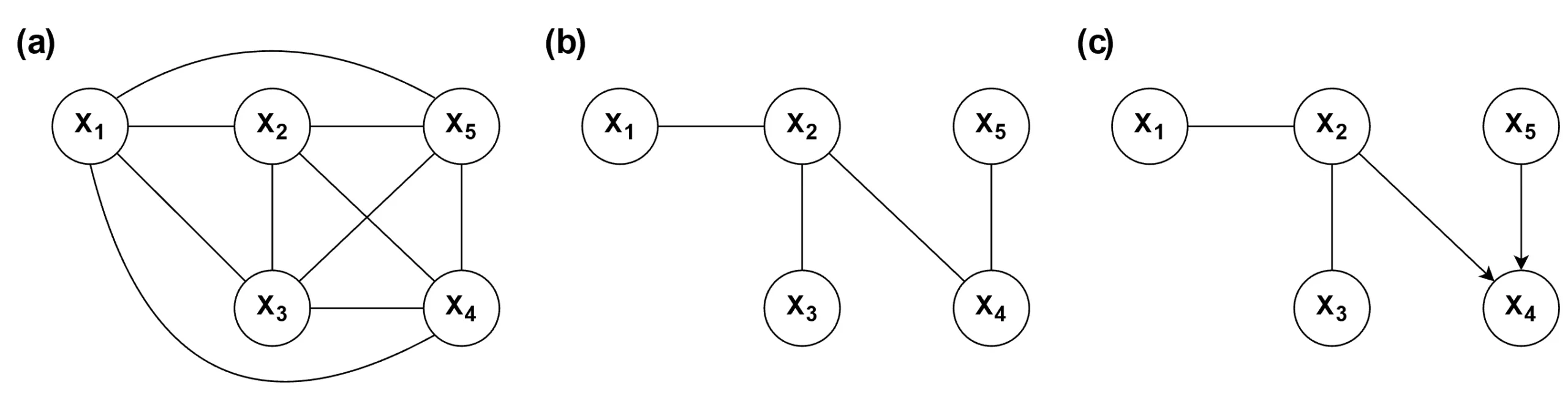
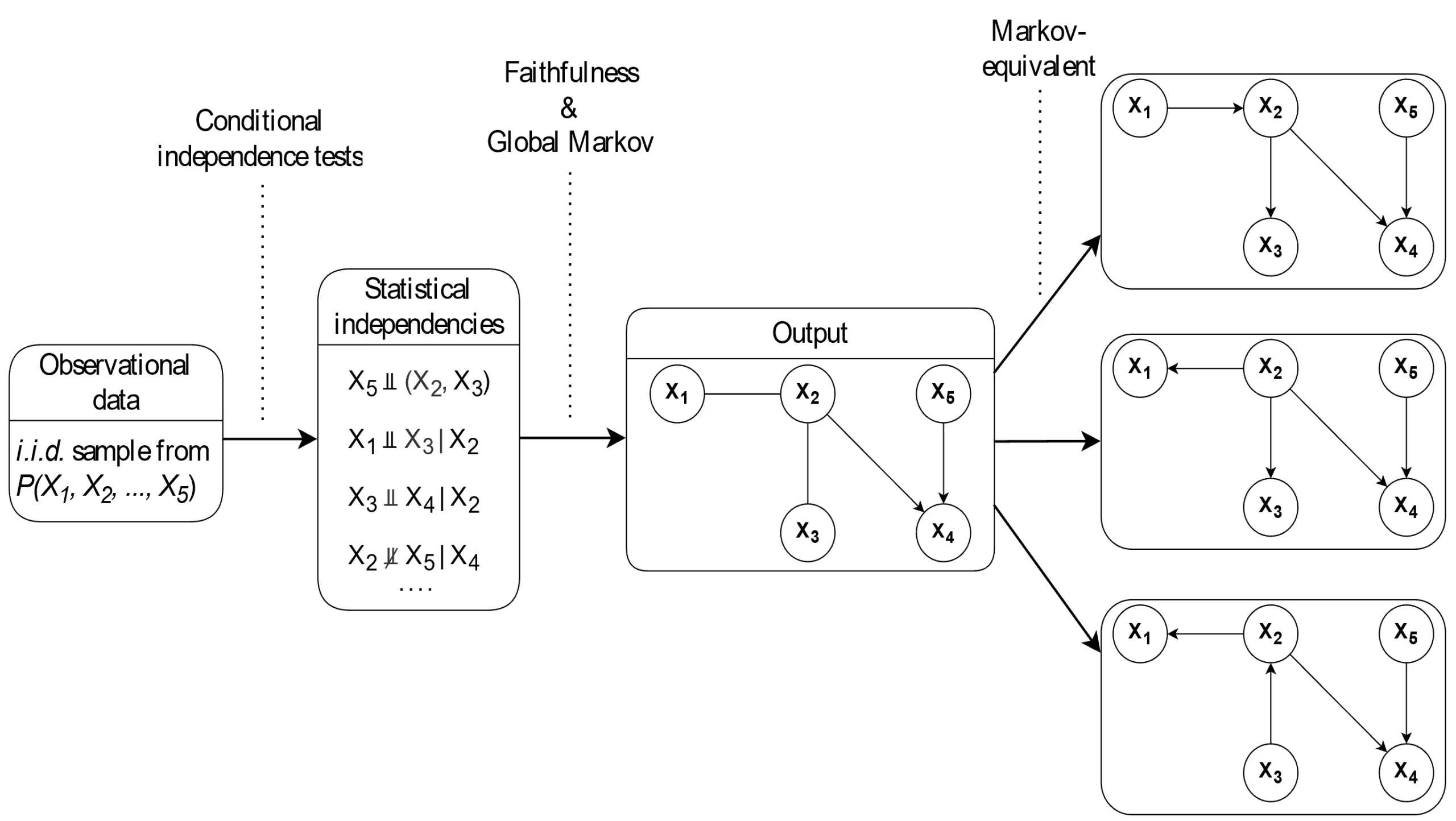
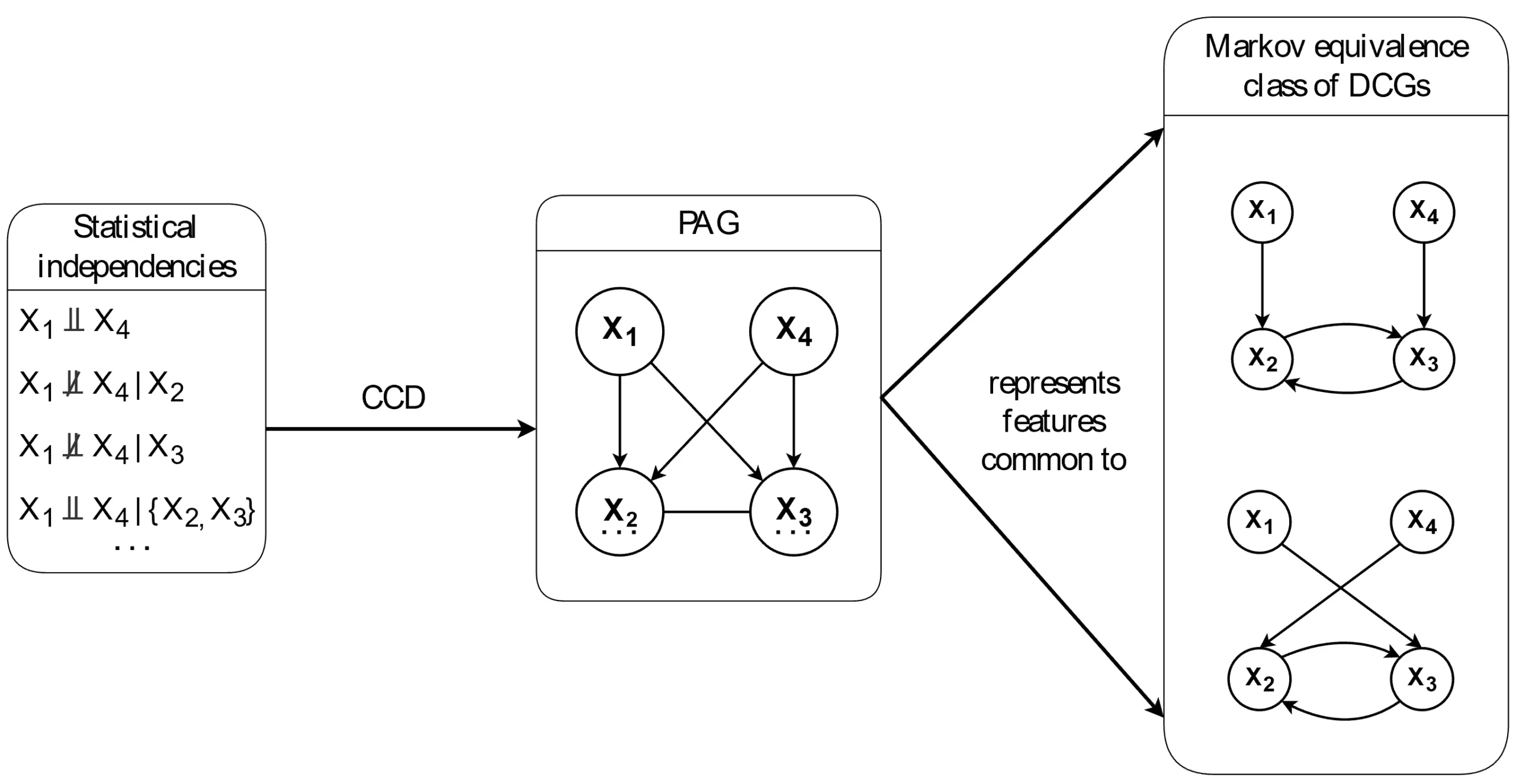
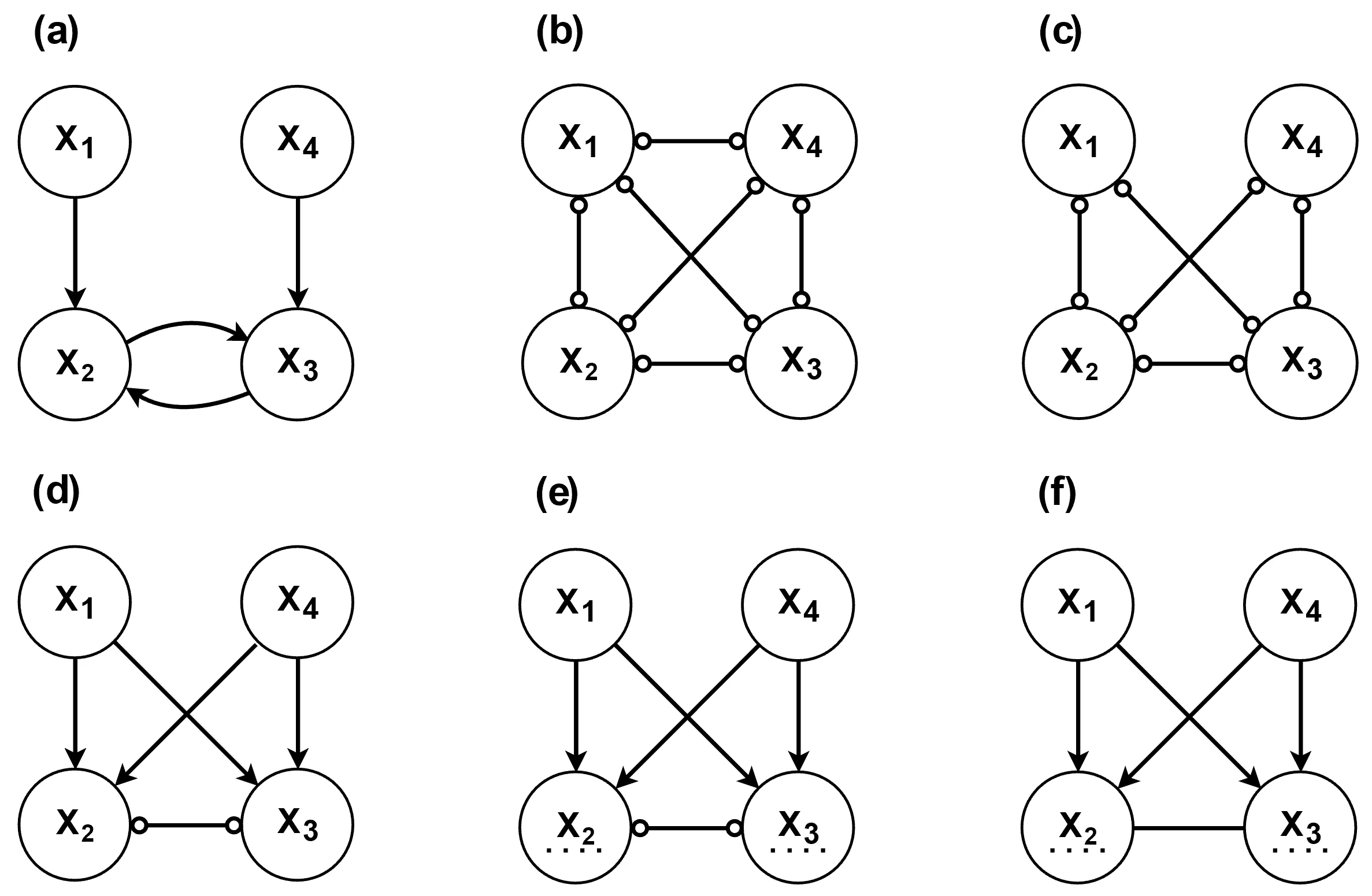
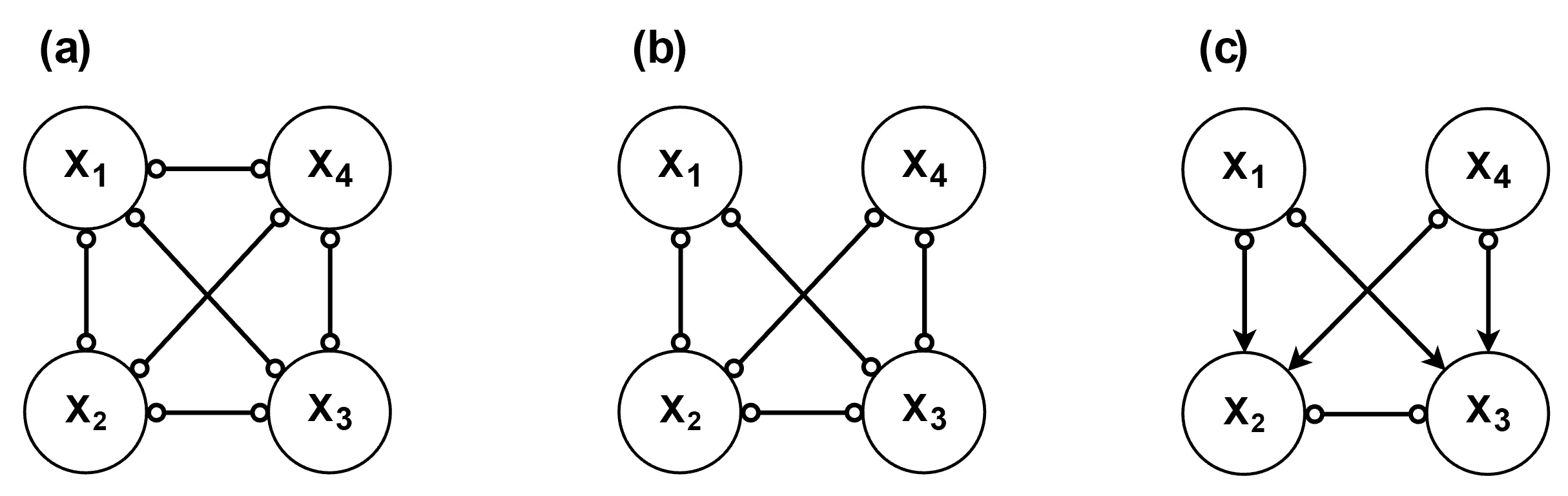
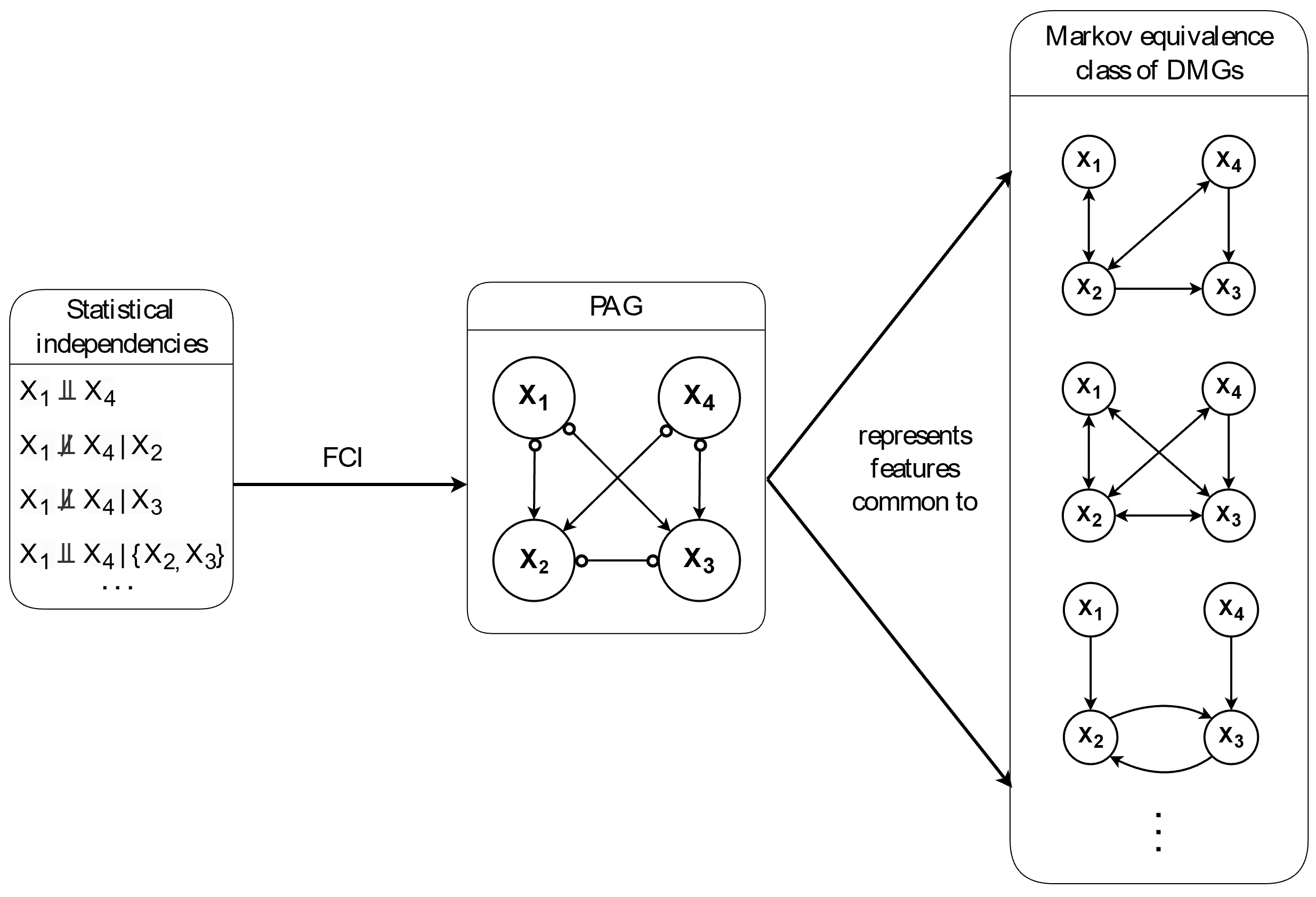
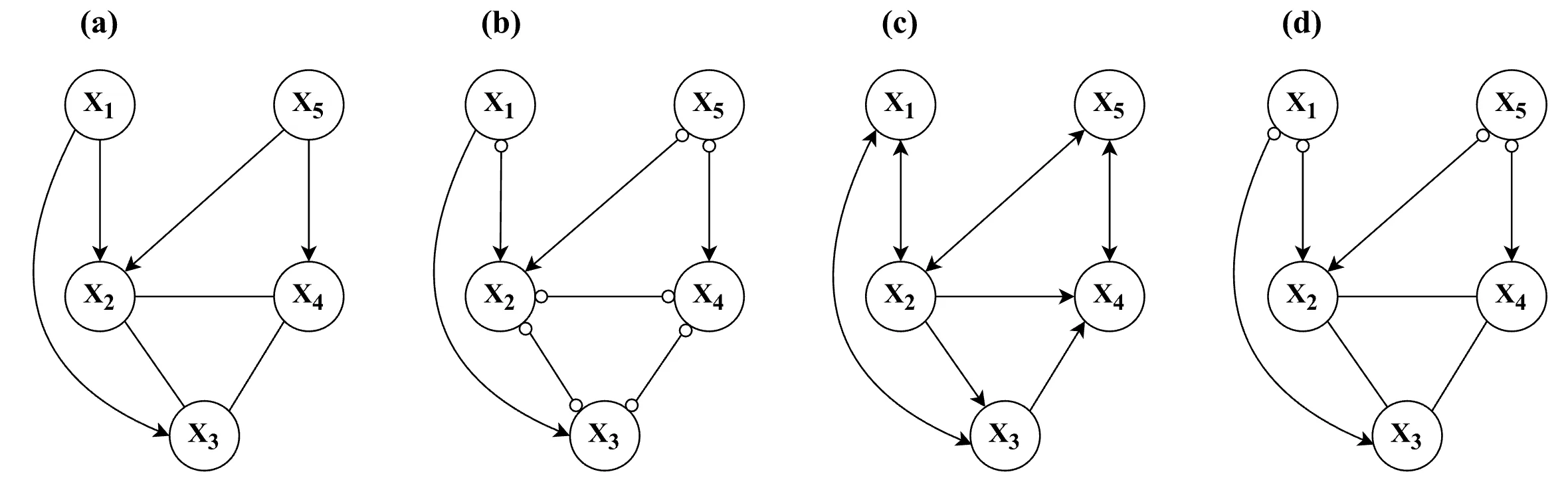
 ) in the graph. As a result, the Markov equivalence class tends to be relatively large.
) in the graph. As a result, the Markov equivalence class tends to be relatively large.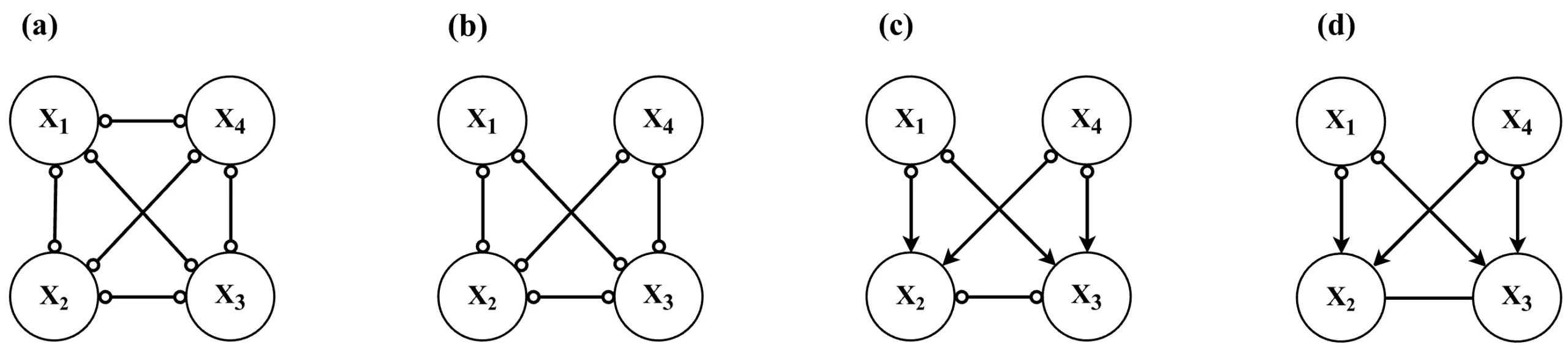
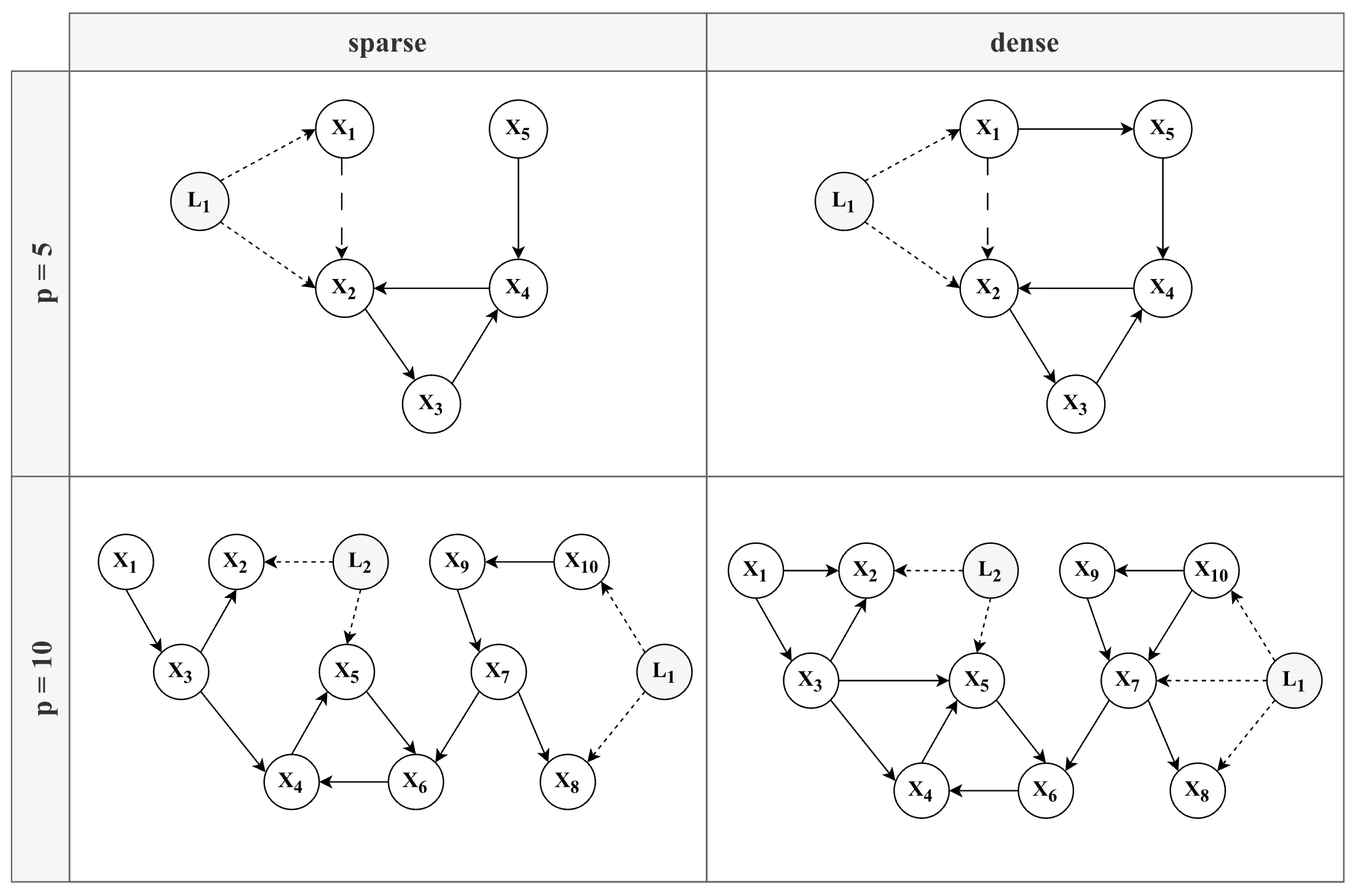
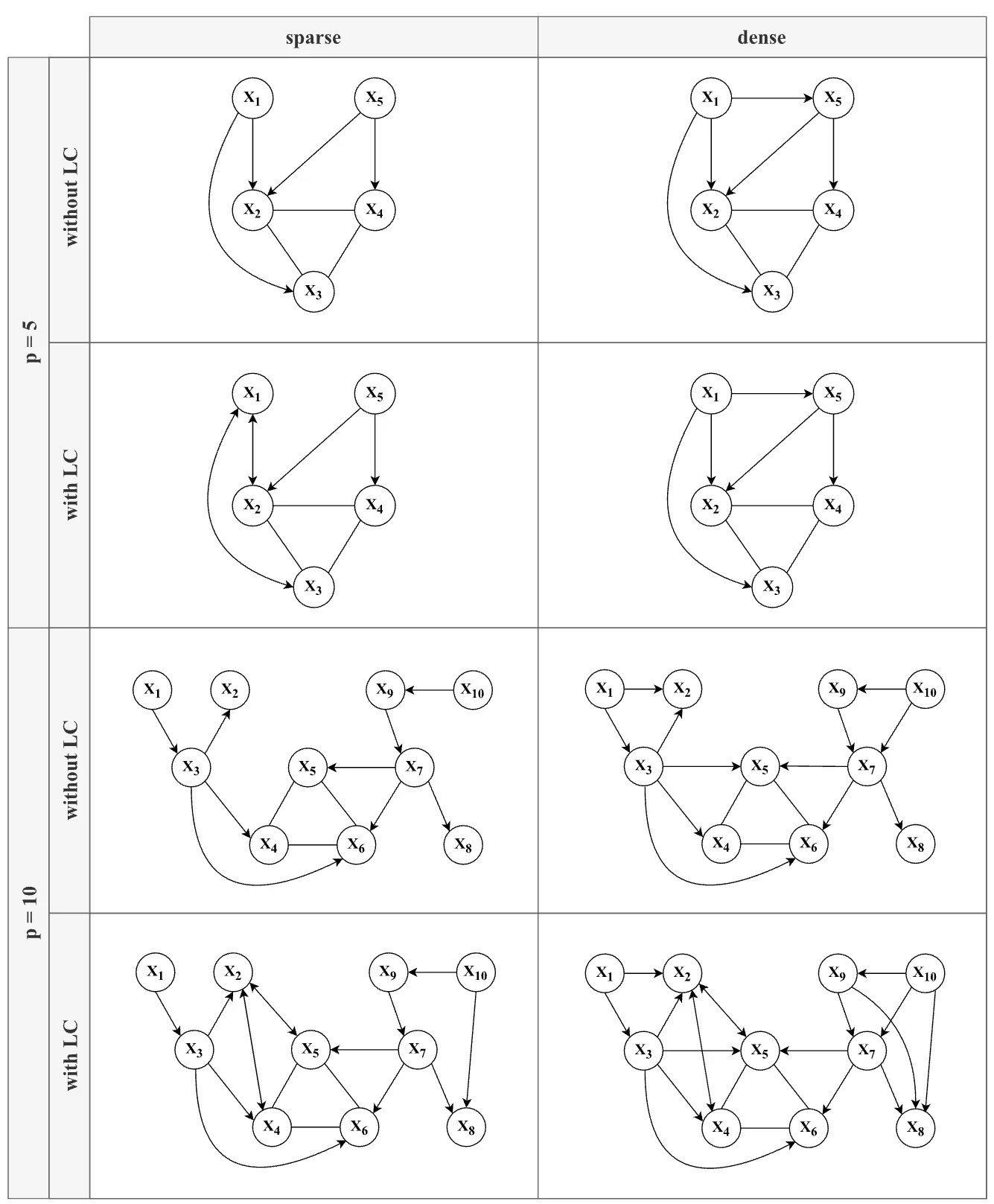
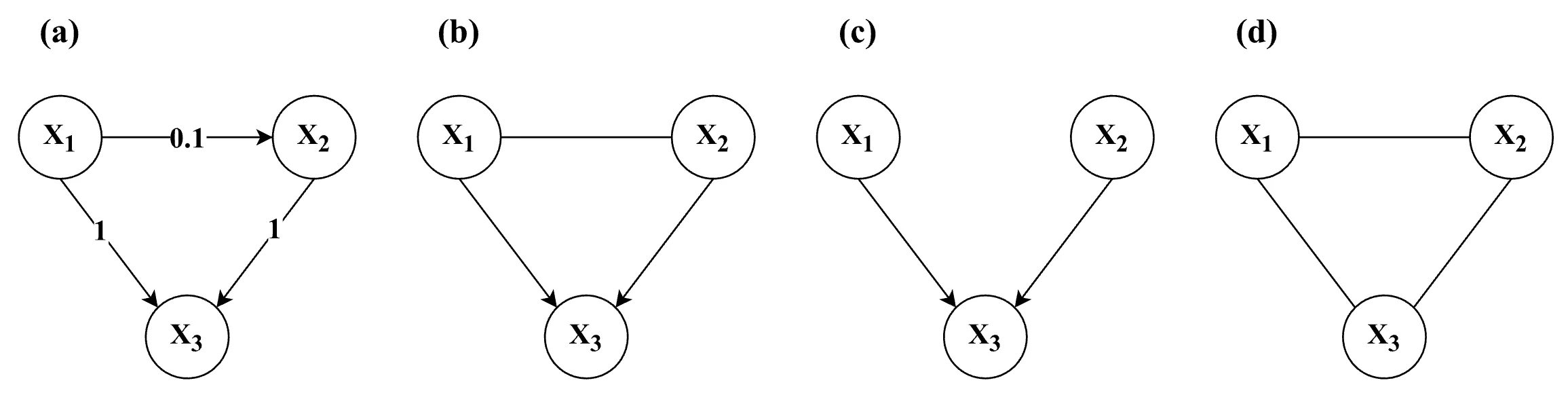
 simulation design, with each combination of factors (except for N) yielding a unique model structure. Note that the edge between X1 and X2 (long-dashed line — —) in the 5-variable models (top row) is only present in the conditions without a latent confounder.
simulation design, with each combination of factors (except for N) yielding a unique model structure. Note that the edge between X1 and X2 (long-dashed line — —) in the 5-variable models (top row) is only present in the conditions without a latent confounder.